Direct to Consumer Genetics of Rheumatoid Arthritis (RA)
RA Genetics 101: Glass half-full or half-empty? Part 2
In part 2 of this guest series, I present results of my own genotyping analysis provided by the direct-to-consumer (DTC) genetic testing company 23andMe, specifically with respect to the estimated lifetime risk of acquiring RA. To provide a somewhat balanced perspective, I also show the corresponding results of lifetime risk of RA for my sister, daughter, and 1st cousin, all of whom have been genotyped by 23andMe. Part 1 of this RA genetic series is available here.
Overview of genotyping for RA risk with 23andMe
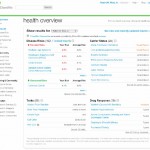 First, in the figure below I present an overview of how 23andMe genotyping results are displayed to the customer.
First, in the figure below I present an overview of how 23andMe genotyping results are displayed to the customer.
Results are divided into four basic categories: Disease Risks, Carrier Status, Traits, and Drug Response.
Others have written about DNA testing to research health disease ancestry or the general topic of using genetic testing with 23andMe; however I will restrict these comments to the first category, Disease Risks. This category is itself divided further into three sub-categories, Elevated Risk, Decreased Risk, and Typical Risk. The full set of results under each sub-category is made available to the customer by mouse-clicking on Disease Risks. In my case, results for RA are categorized (presently; more on this aspect later) under the Decreased Risk category. All other disease risks, while interesting, will not be discussed for purpose of this series. (Note: to those familiar with 23andMe genotyping, my data reflect results obtained from the V2 chip; the newer V3 chip provides updated information.) Currently, 23andMe and most other DTC genetic companies discuss genetic risk in terms of “average lifetime risk” (further discussion on the term “risk”).
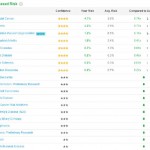 The panel summarizing Decreased Risk according to my genotyping results is shown on the right. Five stars indicates corresponding results that are backed by considerable established research investigations (references are provided). My results suggest that I have a 0.41X risk of developing RA in my lifespan, or about two and one-half times less likely than average.
The panel summarizing Decreased Risk according to my genotyping results is shown on the right. Five stars indicates corresponding results that are backed by considerable established research investigations (references are provided). My results suggest that I have a 0.41X risk of developing RA in my lifespan, or about two and one-half times less likely than average.
Clicking on Rheumatoid Arthritis under the heading Decreased Risks gives the following topic entry page (top half shown). 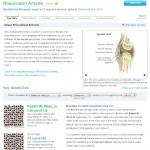 Tabs at the top of the figure allow migration to additional pages of information relevant to RA risk. Of those shown, the one I use most often is “Technical Report”; below I’ll provide specific information from it on my RA risk. Meanwhile, a glance at the panel in this figure titled Your Genetic Data shows that my average lifetime risk of RA is presently 0.98 (rounded up to 1.0 in the preceding figure), versus that of the average population risk (European mainly) of 2.4. Again, this is another representation suggesting I am at about two and one-half times less likely to develop RA over my lifespan than an average individual of European descent. Explaining how best to interpret this single piece of information would require a full post itself, and for now I will defer to a later post in this series. Next, however, I want to focus on the specific genetic findings used to establish this particular risk value for RA in my genotyping results.
Tabs at the top of the figure allow migration to additional pages of information relevant to RA risk. Of those shown, the one I use most often is “Technical Report”; below I’ll provide specific information from it on my RA risk. Meanwhile, a glance at the panel in this figure titled Your Genetic Data shows that my average lifetime risk of RA is presently 0.98 (rounded up to 1.0 in the preceding figure), versus that of the average population risk (European mainly) of 2.4. Again, this is another representation suggesting I am at about two and one-half times less likely to develop RA over my lifespan than an average individual of European descent. Explaining how best to interpret this single piece of information would require a full post itself, and for now I will defer to a later post in this series. Next, however, I want to focus on the specific genetic findings used to establish this particular risk value for RA in my genotyping results.
How risk of RA is established with genetic results
Genetic results from 23andMe are illustrated in two separate ways, first by a summary figure and second by individual risk gene variant (or “allele”) tables. The term “variant” implies more than one version of nucleotide base at a given location in the corresponding gene or locus. Such variants or alleles are often affiliated with different outcomes regarding both risk for developing RA as well as potential, specific disease manifestations once acquired.
First, the summary figure representing my genotyping results for RA is below. This figure shows 9 distinct genetic variants presently tested by 23andMe which are known to impact risk of developing RA from a particular type of research study known as “genome-wide association study” or GWAS. The details are explained in the adjacent legend. One of the nine variants, PADI4, is found only in Japanese populations, reducing the relevant number of genetic variants linked to RA for Europeans like myself to eight, presently. Of those 8, I happen to carry 7 genetic variants placing me at decreased risk of RA, versus one variant placing me at increased risk. The variants are not of equal weight regarding the amount of lifetime risk imposed on the holder; some have much more relevance to developing RA than others (discussed in the next post). Nevertheless, the overwhelming conclusion from these results is that my genetic profile suggests I am less likely than average to develop RA within my lifespan.
By comparison, the results for three female members of my family, all of whom are at increased risk of RA solely due to gender, are shown below. Although all are also clearly at increased genetic risk of RA with respect to me regarding the 8 relevant genetic variants tested by 23andMe, indicated by the red bars (involving both number of red bars as well as height) in the corresponding figures, none has been diagnosed with RA, while only one is above average age of onset (usually > 40 years old) and shows any possible symptoms (cousin).
Of course, everyone knows that phenotype (disease onset and manifestations) is a product of genes X environment (GxE), and that the relationship between these two variables with respect to any given disease is unique. In the specific case of RA, the contribution of heredity to disease risk is shown in the figure below, and indicates that a genetic contribution to RA disease risk falls between one-half and two-thirds of the total risk, with environmental contributions (e.g. in the case of RA, smoking) accounting for the remaining one-third to one-half.
Greater detail regarding the results presented above is provided in table form by 23andMe, shown in the nine distinct but related tables in the figure (below). Each section presents a distinct gene or region or locus having a unique and quasi-independent effect on risk of RA. Where a specific gene is identifiable, as in the case of “PTPN22,” a biological target of RA for therapeutic purposes is available. By contrast, where a specific gene is not identifiable in affiliation with the corresponding genetic variant, as in the case of “HLA region”, the variant is currently useful for diagnostic but not therapeutic purposes. Importantly, the exact genetic variant involved at each gene/locus is provided, indicated as an “rs” number (rs = “reference SNP”; SNP = single nucleotide polymorphism) which can be used for investigative purposes at other online genetic analysis programs such as SNPedia as well as while exploring the original literature. So, for example, the genetic variant responsible for RA risk at the HLA region is designated rs6457617. Two other bits of information are provided for each variant: Genotype, discussed later, and Adjusted Odds Ratio, where a number lower than 0 indicates decreased risk (proportionately) while a higher number indicates increased risk. The numbers in this table correspond to those in the respective summary figure above for me.
 As stated in part 1 of this series, the above information is useful mostly as a foot-in-the-door approach to a discussion of RA genetics. The next post will describe in more detail the specific genes, variants, biological pathways and functions, and most importantly, describe present limitations with genetic approaches to diagnosing and treating RA. The reader will rapidly come to appreciate both the promises and perils of investing precious resources (time and money) in this particular approach to understanding and treating human disease conditions.
As stated in part 1 of this series, the above information is useful mostly as a foot-in-the-door approach to a discussion of RA genetics. The next post will describe in more detail the specific genes, variants, biological pathways and functions, and most importantly, describe present limitations with genetic approaches to diagnosing and treating RA. The reader will rapidly come to appreciate both the promises and perils of investing precious resources (time and money) in this particular approach to understanding and treating human disease conditions.
Note: I hope you’re enjoying this guest series on genetics and RA by Bob West. Stay tuned for more!





This is so interesting, thank you for taking the time to do this for us. My mother has RA as well as my older sister and myself. I seem to have it the worst, many more joints involved. I was a smoker for many years, wonder if that was the difference? I quit two years ago before I was diagnosed, started again for a few months earlier this year, and quit again back in March. My pain was worse while I was smoking. Sure hope they come up with a cure soon, I have a daughter and two grand daughters. Don’t want them to deal with this misery.
Becky,
Sorry to hear about your FH of RA. It does not surprise me at all, however, that your symptoms are worse than your other family members after years of smoking, the #1 environmental cause of RA incidence as well as disease severity. Kelly can probably refer you to her prior posts on this topic; it’s certainly an issue that all #Rheum Docs should be fully aware of. I’m not an M.D., but with a FH of RA your kids are at increased risk. Above all, they should be alerted to the increased complications of the Genes X Environment combo of RA & smoking.
-Bob
Thank you for the reply. If I am understanding correctly, smoking caused my RA to be more involved(more joints affected) or does it cause RA to be more painful ? And should the severity and pain decrease once you stop or is the damage permanent?
I do feel my pain was worse when I started again and seemed to improve when I quit. But when you deal with a constant level of pain sometimes it is hard to tell how much… I am angry at myself for causing this to happen but I will take from this that I can warn my daughter and grand daughters to stay away from smoking.
Dear Becky,
I hope it’s ok if I comment. First of all, I don’t think anyone could know for sure what the future will be like with RA flares. There are studies that say smoking makes RA worse and makes treatment less effective, so quitting could help people who already have RA, like you said you probably noticed. I am so sorry you are dealing with these feelings on top of the stupid disease. Please don’t be too hard on yourself.
Also, you can’t know for sure that your smoking was the only trigger for your RA to start. There is more than 1 trigger since there are folks who have RA who have not smoked. True, the message is this teaches us is: we should not smoke. But no one would say this is all your own fault, since there are genetic factors that were beyond your control. How many smoke and do not get RA? Back when you started, you had no idea that you could have been contributing to this.
Kelly, thank you for your concern. I’ll work through it but yes, it is hard to face that you have done something that could have caused you to suffer for the rest of your life. It is tough enough to deal with the lack of understanding from those around you.
One thing I think it means though is that it could be one of the reasons that the medication is not helping me like my Dr. thinks it should. Another reason is that my Dr. is fairly new and is treating by the book, glad he is gone and I get a new one next time I go. Keeping my fingers crossed(mentally cause I really can’t do that now!)that I get a Dr. that is more flexible.
Becky,
Glad Kelly spoke first since her knowledge of RA is obviously much broader and deeper than mine, but I’m also confident her answer to your question is right on the money! I thought what I might do to contribute is to provide a key reference on the issue of RA genes X environment. The attached link is to a paper entitled “Environmental influences on risk for rheumatoid arthritis” by Elizabeth Karlson, a key investigator on RA, and this paper is perhaps one of the seminal reviews on this topic: http://www.ncbi.nlm.nih.gov/pubmed/19318947. Fortunately, it’s also freely available from PubMed Central (tab on upper right when you open the link). Although this review is now 2 years old, environmental determinants is one of the slowest moving areas of RA research, similar to most other areas of chronic disease investigation. Thus, you may still be left with unresolved questions about “what might be” and “what might have been”.
Thank you for the link, very interesting that it mentions “the highest risk of seropositive RA was in heavy smokers who carried two copies of the shared epitope with evidence for multiplicative interaction.” I am seronegative. I do wonder however since I have only been tested twice and both times was after heavy does of steroids, if the test was accurate. My RA was verified by ultrasound. I am also 55 so onset was at an older age.
I don’t want to obsess with this. However I do feel I owe it to myself to research and try to understand what is going on with my body. And again what I learn might help my family to avoid having this happen to them.
Thanks again for taking the time write all of this out for the benefit of others (especially those of us with non-science backgrounds!). How amazing that you can get such detailed genetic information about yourself! I’m very curious what my results would show. Interesting about the presumed contribution of genetic vs. environmental factors influencing RA disease manifestation…I wonder what other environmental factors might play a part besides smoking? I look forward to your subsequent posts in this series!
Thanks Dana. Smoking is the really big environmental determinant that affects RA, both increased incidence and increased severity once initiated. At some point I will try to provide refs. (I have here, just too tired to check right now :-/ ). Unfortunately, few other environmental factors have been convincingly shown to be associated with RA. This is a big black box area for future investigation, and IMO will be more difficult to sort out than the genetic contributions to RA.
-Bob
Dana,
I’ve now provided a key reference on environmental factors thought to be associated with RA. See the reply to Becky above, by both Kelly and me.
-Bob
Where are the references posted? Very interesting, Thanks!
Debra,
Sorry, the link was embedded in my 2nd reply to #1-Becky. If you are colorblind (not out of the question, one of my two daughters is) you likely would not see it, since (likely) red and not underlined. I’m posting the link again here: http://www.ncbi.nlm.nih.gov/pubmed/19318947.
-Bob
Pingback: More Genetic Connections with RA « Living with Rheumatoid Arthritis
Hi Kelly,
This is an old article, and many more SNPs have been associated with the risk of developing RA. If you haven’t already, you might consider uploading your raw data from 23 and me to https://promethease.com
It will compare your DNA against scientific findings. It is fascinating, and you can come to your own conclusions by reading the relevant scientific studies.
BTW, I loved your book!